Acute promyelocytic leukemia with PML/RARA (bcr1, bcr2 and bcr3) transcripts in a pediatric patient
- Authors:
- Published online on: January 22, 2024 https://doi.org/10.3892/ol.2024.14246
- Article Number: 114
-
Copyright: © Santana-Hernández et al. This is an open access article distributed under the terms of Creative Commons Attribution License.
Abstract
Introduction
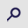
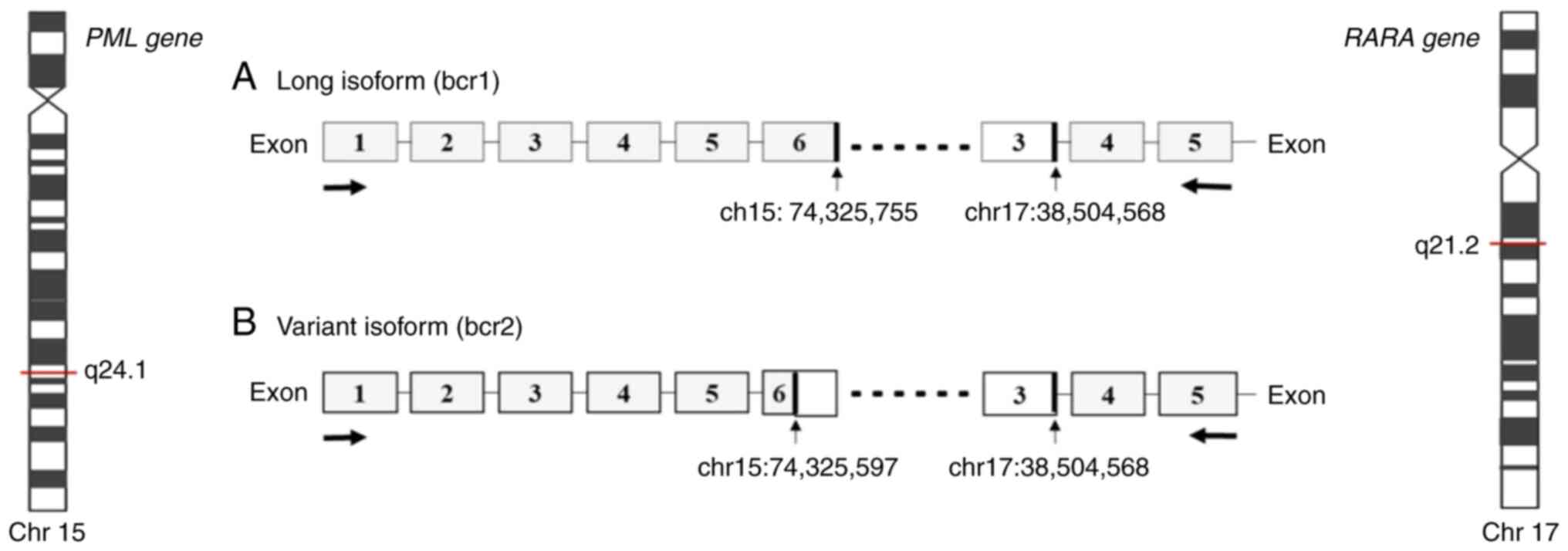
Acute promyelocytic leukemia (APL) represents ~10% of all childhood acute myeloid leukemia (AML) cases (1,2). Cytomorphologically, APL is classified as hypergranular (or typical M3) or as hypogranular (or variant M3V) (3). APL presents a balanced reciprocal translocation t(15;17)(q24.1;q21.2), involving the promyelocytic leukemia (PML) gene on chromosome 15 and the retinoic acid receptor α (RARA) gene on chromosome 17 (4,5). There are three distinct PML breakpoint cluster regions, namely bcr1, bcr2 and bcr3, that generate a PML/RARA fusion gene of varying mRNA lengths. The bcr1 transcript results from a break in intron 6 of PML with exon 3 of RARA; transcript bcr2 results from a break in exon 6 of PML with exon 3 of RARA; transcript bcr3 results from a break in intron 3 of PML with exon 3 of RARA (6). The frequency of the different breakpoints is dependent principally on ethnicity and geographical location as the bcr1 transcript is more frequent among individuals of Latin origin (3). The aforementioned transcripts have been detected using reverse transcription-quantitative PCR (RT-qPCR) (7,8). The PML/RARA fusion gene generates a PML/RARA oncoprotein that blocks myeloid precursor differentiation, leading to the accumulation of abnormal promyelocytes in the bone marrow (9,10). The PML domain of the PML/RARA oncoprotein can affect the senescence pathway, facilitating the acquisition of mutations that drive the development of leukemia (9–11). Investigations involving children and adults have provided information on the typical and atypical transcripts of PML/RARA. Typical transcripts are the most common (identified in 90–95% of cases) and have three breakpoints in intron 6, exon 6 and intron 3 of the PML gene and only one in intron 2 of the RARA gene. Atypical transcripts have breakpoints in intron 4, exon 6 or exon 7 of the PML gene, and in intron 3 of the RARA gene. However, the biological function of atypical transcripts in the initiation and evolution of APL have yet to be defined (11). The present study aimed to describe a novel pediatric case that presents atypical bcr1, bcr2 and bcr3 transcripts, as well as the clinical and molecular aspects present in this APL case. To the best of our knowledge, the present study is the first to identify the three aforementioned atypical PML/RARA transcripts.
Materials and methods
Case report
A 12-year-old female patient was admitted in March 2018 to the Civil Hospital of Guadalajara (Guadalajara, Mexico) with gingival bleeding, hyperplasia, petechiae, ecchymosis, paleness and traces of bleeding in the oral cavity, with no lymph node enlargement, hepatomegaly or splenomegaly. Laboratory analyses revealed the following: Hemoglobin levels, 10.8 g/dl (normal range, 12–16 g/dl); leucocytes, 2,670/µl (normal range, 5,000-10,000/µl); platelets, 8,000/µl (normal range, 150,000-400,000/µl); prothrombin time, 13.1 sec (normal range, 9–13 sec); activated partial thromboplastin time, 25.7 sec (normal range, 26–40 sec); and D-Dimer levels, >1,500 ng/ml (normal range, 340–729 ng/ml). Bone marrow aspiration revealed that 98% of nucleated cells were replaced by myeloblasts, that hypergranular promyelocytes were densely packed, bright-pink, reddish-blue or dark-purple granules, and that there were numerous Auer rods. Immunophenotyping revealed a population of 91% of promyeloblasts, which was CD13+, human leukocyte antigen (HLA)-DR−, CD38+, CD117+ and CD45+. All the aforementioned data were compatible with Acute Myelomonocytic Leukemia or French-American-British (FAB) M3 classification (12). The patient was staged at intermediate risk according to the PETHEMA APL 2012 protocol proposed by Spanish Society of hematology and hemotherapy (13). The patient achieved remission on day 50 after receiving consolidation therapy with three chemotherapy cycles, which was maintained for 2 years according to the protocol. The patient completed treatment 4 years ago (2018–2022). The patient has a good prognosis and continues to be followed up.
Ethical considerations
The present study was submitted and accepted by The Research Committee and The Research Ethics Committee of The Civil Hospital of Guadalajara (approval no. 00116). Bone marrow aspirates from the patient and reference (wild-type control) were obtained prior to treatment. Written informed consent was obtained from the parents and the institutional review boards approved the use of excess diagnostic material for research purposes. These studies were conducted in accordance with the Declaration of Helsinki.
Karyotyping and fluorescence in situ hybridization (FISH) analysis
A bone marrow sample of the patient was obtained and G-banding karyotyping was performed. FISH analysis was then performed for the detection of the translocation, t(15;17)(q24.1;q21.2). Cells were dropped onto glass slides to perform the FISH assays, which were conducted following the manufacturer's recommendations. Images were captured using an AXIO ImagerMI (Zeiss AG) microscope, and the images were analyzed using ISIS software (MetaSystems). A total of 200 interphase cells were reviewed in each slide. The PML and RARA genes were analyzed using a Vysis LSI PML/RARA Dual Color probe, Dual Fusion Translocation Probe (cat. no. 05J70-001; Abbott Molecular, Inc.). For the dual-color probe, cells with 1 orange, 1 green and 2 fusion signals were considered positive for the PML/RARA fusion.
RNA isolation and RT-PCR
RNA was isolated from lymphocytes of the bone marrow using the method of TRIzol™ (cat. no. 15596026; Thermo Fisher Scientific, Inc.) proposed in the study by Rio et al (14). The Applied Biosystems™ High-Capacity cDNA RT kit (cat. no. 4368813; Applied Biosystems; Thermo Fisher Scientific, Inc.) was used for cDNA synthesis. The reaction included 4 µl 10× RT buffer, 4 µl 10× RT random primers, 1.8 µl 25× dNTP Mix (100 mM), 50 U/µl MultiScribe® Reverse Transcriptase and 1 µg RNA, and was conducted using an Applied Biosystems ProFlex PCR system (Applied Biosystems; Thermo Fisher Scientific, Inc.) under the following conditions: 10 min at 25°C, 2 h at 37°C and 5 min at 85°C.
Detection of PML-RARA rearrangements
cDNA was analyzed using the HemaVision-28N Multiplex RT-qPCR kit in the Applied Biosystems ProFlex PCR system (Applied Biosystems; Thermo Fisher Scientific, Inc.) and the Qualitative HemaVision-28Q RT-qPCR kit (cat. no. HV01-28Q; DNA Diagnostic A/S) in the Rotor-Gene® Q system (cat. no. 1070452EN; Qiagen, Inc.). These methodologies can identify 28 chromosomal translocations and >145 gene breakpoints associated with leukemia. The procedures were performed according to the manufacturer's protocols. Duplicate analysis was considered in the methodology using quality and negative controls from the kit.
An adapted RT-qPCR protocol from the study by Gabert et al (15) was used to identify the PML/RARA fusion gene. The primers used were as follows: i) ENF903 (bcr1 forward, 5′-TCTTCCTGCCCAACAGCAA-3′; 19 bp); ii) ENF906 (bcr2 forward, 5′-ACCTGGATGGACCGCCTAG-3′; 19 bp); iii) ENF905 (bcr3 forward, 5′-CCGATGGCTTCGACGAGTT-3′; 19 bp); iv) ENR962 (bcr1-3 reverse, 5′-GCTTGTAGATGCGGGGTAGAG-3′, 21 bp); and v) ENP942 (probe, 5′ FAM-AGTGCCCAGCCCTCCCTCGC-BHQ-1 3′, 20 bp). The RT-qPCR was conducted using the following reagents: 12.5 µl TaqMan Gene Expression Master Mix (cat. no. 4369016; Applied Biosystems; Thermo Fisher Scientific, Inc.), 1.2 µl forward and reverse oligonucleotides, 0.5 µl probe, 8.6 µl nuclease-free water and 1 µl cDNA. The samples were placed in a 96-well plate and analyzed using the 7900 HT Fast Real-Time PCR system (Applied Biosystems; Thermo Fisher Scientific, Inc.) with the SDS version 2.4 program (Thermo Fisher Scientific, Inc.). The thermocycling conditions were as follows: 50°C for 2 min, 95°C for 10 min, 95°C for 15 sec, and 60°C for 1 min (40 cycles). To confirm the expression of PML/RARA transcripts, the NB4 human cell line [derived from the leukemic cells of a relapsed acute promyelocytic leukemia (M3) patient and carrying the t(15;17) translocation; CVCL_0005; Cellosaurus, https://www.cellosaurus.org/CVCL_0005] was donated by St. Jude Children's Research Hospital (Memphis, USA), and validated samples positive for bcr1, bcr2 and bcr3 variants were used as positive controls. The NB4 cells were cultured in RPMI 1640 medium (Gibco; Thermo Fisher Scientific, Inc.; cat. no. 11875093) with 15% fetal bovine serum (Gibco; Thermo Fisher Scientific, Inc.; cat. no. A4766801), 2 mmol/l L-glutamine (Gibco; Thermo Fisher Scientific, Inc.; cat. no. 25030081), 1X antibiotic (Gibco; Thermo Fisher Scientific, Inc.; cat. no. 15240062) and incubated in a humidified incubator at 37°C with 5% CO2. The RNA isolation and RT-qPCR protocol for the NB4 cells was the same as aforementioned. Non-template controls (nuclease-free water) were also included in each experiment. The quantification of relative expression of the transcripts was calculated using the 2−ΔΔCq method, with β-glucuronidase was as the housekeeping gene.
Genomic analysis
cDNA obtained using RT was analyzed to detect simultaneous PML/RARA transcripts and myeloid leukemia-associated genes using the Illumina® AmpliSeq RNA Myeloid Panel (cat. no. 20024478; Illumina, Inc.). The procedure was performed by Illumina Inc., and each RT reaction required 10–100 ng total RNA.
Expression microarray
A wild-type sample (from a healthy 10-year-old patient) and the APL case were used for the expression microarray. RNA reconstituted in UltraPure™ DEPC-Treated Water (cat. no. 750023; Invitrogen; Thermo Fisher Scientific, Inc.) was quantified and examined for RNA quality using a NanoDrop 2000™ spectrophotometer (cat. no. ND-2000; Thermo Fisher Scientific, Inc.). The A260/A280 and A260/A230 ratios between 1.8 and 2.2 were used to determine the RNA quality. RNA integrity was evaluated using a 15% agarose gel stained with Gel Red (cat. no. 41003; Biotium, Inc.), visualized using a FirstLight® Uniform UV Illuminator (model, LM-20; single intensity; 302/365 nm UV; filter size, 20×20-cm; 230 V; cat. no. 95-0449-02).
To detect transcripts, the human Clariom™ D Assay (cat. no. 902922; Applied Biosystems; Thermo Fisher Scientific, Inc.) was used. The Applied GeneChip System 3000Dx v.2 included the GeneChip® Hybridization Oven 645 with GeneChip® Fluidics Station 450 Dx v.2 and workstation. Array images were acquired using a GeneChip® Scanner 3000Dx v.2 with AutoLoaderDx and Affymetrix Molecular Diagnostic Software (Affymetrix; Thermo Fisher Scientific, Inc.).
Data analysis and functional classification of differentially expressed genes
All data were captured using Applied Biosystems Transcriptome Analysis Console (TAC) software (version 4.0.2; Thermo Fisher Scientific, Inc.) and microarray data were deposited in the Gene Expression Omnibus (GEO) database following the Minimum Information about a Microarray Experiment (MIAME) and Minimum Information about a Next-generation Sequencing Experiment (MINSEQE) guidelines (https://www.ncbi.nlm.nih.gov/geo/). Finally, to identify candidate differentially expressed genes, the microarray data were analyzed using the online Database for Annotation, Visualization and Integrated Discovery (DAVID) Bioinformatics Resources (version 6.8; http://david-d.ncifcrf.gov/). A fold change ±1.5 and P<0.05 were considered to indicate a statistically significant difference in expression. However, only those genes with a fold change > ±10.0 (the highest selection) were selected.
Results
Conventional cytogenetics
Karyotyping analysis of the patient revealed 46,XX,t(15;17)(q24;q21)[20] and the FISH result was nuc ish(PML,RARA)x3(PML,RARA)x2[148/200](PML,RARA)x2[52/200], with [n/n] representing the number of cells counted with the alteration out of the total (Fig. S1).
HemaVision RT-qPCR test
All three transcripts were identified using the HemaVision-28N Multiplex RT-PCR kit, with a size of 353 bp for bcr1 (PMLex6-RARAex3), 97–350 bp for bcr2 (PMLδex6-RARAex3) and 325 bp for bcr3 (PMLex3-RARAex3) (Fig. S2). Furthermore, the atypical transcripts were corroborated using the Qualitative HemaVision-28Q RT-qPCR assay. The Cq values were as follows: i) 29.83 (bcr1, PMLex6a-RARAex3); ii) 30.59 (bcr2, PMLex5-RARAex3); and iii) 30.12 (bcr3, PMLex3-RARAex3). It should be noted that the breakpoints reported are dependent on the primer design in each kit.
RT-qPCR data
The measurable residual disease was calculated at diagnosis through the relative expression quantification of the simultaneous transcripts using the Livak method (16). The results were 100% for bcr1 (Cq, 28.931), 38.5% for bcr2 (Cq, 30.08) and 0.83% for bcr3 (Cq, 35.86). After the first month of treatment, expression of the three PML/RARA transcripts decreased to 0%.
Genomic analysis of simultaneous transcripts in the patient with APL
Genomic analysis with RNA sequencing. Genomic analysis using the AmpliSeq RNA Myeloid Panel Illumina® revealed two simultaneous transcripts, bcr1 and bcr2 (Fig. 1).
Data analysis of differentially expressed genes
First, the microarray data were deposited in the GEO database (accession no. GSE205372; http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE205372), following the MIAME and MINSEQE guidelines. Then, the overall pattern of gene expression in the microarray was assessed. Using TAC Software version 4.0.2, a cut-off fold change value of ±10.00 was set to generate a reduced gene list since only 1 case was analyzed. As a result, 613 differentially expressed genes were identified (139 upregulated and 473 downregulated genes) compared with the control (wild-type). The top upregulated and downregulated genes are listed in Table SI.
Classification of differentially expressed genes in the APL case
Using the data obtained of the 613 differentially expressed genes, two different analyses were performed: i) First, a search was performed for groups of genes arranged by functional similarity and related to APL from the list using the Functional Annotation Clustering setting in DAVID. An enrichment score >2 was deemed a significant enrichment with the highest stringency. The analysis generated several functional clusters of significantly upregulated or downregulated genes (Table I), such as: i) Immune system-related clusters: Immunoglobulin C1-set molecules involved in the immune system, major histocompatibility complex (MHC) class II and the loss of HLA-DR antigen expression; ii) C-type lectin cluster; iii) Src-homology 2 domain (SH2) cluster; and iv) mammalian defensins cluster. Additional clusters unrelated to APL were also found, including graft-vs. -host disease (the patient was previously transfused), Btk motif, AIG1, peptidase SI and Pleckstrin homology domain. The enrichment score is a modified form of the P-value of the exact Fisher test, and the Benjamini value is the adjusted P-value resulting from the Benjamini and Hochberg method.
Table I.Functional annotation clusters obtained by Database for Annotation, Visualization and Integrated Discovery analysis. |
ii) Second, from the list generated and presented in Table SI, only 21 genes with a fold change >20 and 24 genes <-50 were selected that can be associated with clinical and molecular characteristics of APL for future research. The selected genes are part of different signaling pathways, such as for the cell cycle, proliferation, differentiation and adhesion, in addition to MHC and cytokine genes.
Literature review
The present study suggested a possible signaling pathway involving the PML/RARA oncoprotein that leads to cell proliferation or the evasion of apoptosis, based on the microarray analysis (613 up or downregulated genes) and a literature search. The identified genes were selected as ‘key words’ in the literature search in association with characteristic clinical and molecular APL. The following databases were searched: PubMed (https://pubmed.ncbi.nlm.nih.gov/), Kyoto Encyclopedia of Genes and Genomes (https://www.genome.jp/kegg/), Gene-Cards (https://www.genecards.org/), UniProt (https://www.uniprot.org/) and Ensembl (https://www.ensembl.org/index.html).
Discussion
APL is a subtype of AML with a unique molecular appearance distinguished cytogenetically by balanced reciprocal translocation t(15;17) and PML/RARA gene fusion. Furthermore, it is often associated with a complex coagulopathy known as disseminated intravascular coagulation (DIC), resulting in a high hemorrhage rate and thrombosis. Patients with APL present more frequently with severe DIC due to the increased expression of tissue factors and annexin II, which activates fibrinolysis. During the induction period, hemorrhaging is responsible for the majority of deaths (17,18).
Case studies with PML/RARA transcripts are common among adults and uncommon among children (10,19). Generally, one transcript is usually detected in children from common or typical breakpoints and rarely from atypical breakpoints. The bcr1 or bcr3 transcripts are the most frequent (90–95%) and bcr2 is infrequent (11,19–22). Previously, a case bearing two transcripts, bcr1 and bcr2, was detected using the HemaVision RT-qPCR kit, in a pediatric patient with APL detected by immunophenotyping, but with normal results from karyotyping and FISH; however, there was insufficient evidence to define a prognostic factor (23,24). To the best of our knowledge, there have been no cases reporting the presence of the three different transcripts (bcr1, bcr2 and bcr3). In the present study, alternative methods, such as genomic analysis, were applied to identify the breakpoints between the PML and RARA genes and only bcr1 and bcr2 transcripts were detected. Due to the large size of the transcriptome, the RNA sequencing methodology cannot detect fusion genes that are expressed at low levels in the leukemic clones, which may have affected the detection of bcr3 (25).
The effect of the PML/RARA molecular transcripts on APL continues to be controversial as there have been no conclusive results. The reason for the generation of the three bcr1, bcr2 and bcr3 transcripts remains unknown, although it may involve molecular heterogeneity or unidentified secondary alterations. Adaptive advantages provided by such events may contribute to a mutated phenotype during the development of APL (26). To the best of our knowledge, no studies to date mention differences in the oncoprotein expressed depending on the transcript present; thus, it would be of interest to obtain further knowledge of the molecular function of oncoproteins in association with the prognosis of these patients. It can be hypothesized that different PML breakpoints that lead to protein variants could affect the prognosis or therapeutic response; however, this issue remains unsolved (27).
Previous studies have reported that pediatric patients have a >25% frequency of the hypogranular morphological subtype, and a higher frequency of the bcr2 and bcr3 transcripts, compared with adult patients (3,17). In addition, a higher incidence of the bcr1 transcript has been observed in the Latin American population, and the bcr3 transcript is associated with a worse prognosis, the M3v subtype, hyperleukocytosis and a higher frequency of mutations in the Fms-like tyrosine kinase 3 (FLT3) gene. According to the clinical follow-up of the patient, it is suggested that the presence of the three transcripts infers a good prognosis. As the expression of bcr1 and bcr2 inhibits the effects of bcr3, we speculate that there could be environmental influences determining the breakpoint in the PML gene, or there is an additional secondary alteration, as mutations in the FLT3 gene may alter cells to infer an adaptive advantage; however, the present case did not present with FLT3 mutations, or other clinical risk characteristics (17,28).
In the present study, to predict the possible functional interactions of the PML/RARA oncoprotein, an expression microarray analysis was performed. Only gene expression profiles of genes consistently associated with APL were selected, identifying the following clusters of genes: i) Immunoglobulin C1-set molecules involved in the immune system, MHC class II and the loss of HLA-DR antigen expression; ii) C-type lectin; iii) SH2; and iv) mammalian defensins. HLA-DR is a molecule of antigen-presenting cells. The principal function of HLA-DR is to initiate and promote the immune response, and its expression is present in the early stages of the APL disease (29). HLA-DR- and low CD34 expression characterizes malignant promyelocytes. A study by Dunn et al (30) examining the mechanisms of immune evasion indicated an association with the downregulation of HLA-DR antigen expression in tumor cells. The mechanism of immune evasion is an immunoediting process that has been described in transplanted patients with AML (30). The transplanted immune cells exert selective pressure against AML cells that can be recognized immunologically. Tumor clones evolve in response to selective pressure mediated by the immune system and finally escape, leading to resistant clones and relapse. These epigenetic alterations suggest that therapeutic strategies to re-sensitize AML cells to the graft-vs. -leukemia effect may be feasible (29,30). Studies have shown that the expression levels of either CD56, CD34 or FLT3-internal tandem duplication (ITD) markers are associated with a poor patient prognosis (31–33). However, the prognosis of patients with APL expressing CD2, CD4, HLA-DR and FLT3-ITD mutation remains controversial. To the best of our knowledge, pediatric cases have not yet been reported.
The C-type lectin cluster was also found in the RNA analysis of the present study. C-type lectin functions as a recognition molecule in the immune system and has a variety of roles in the defense against pathogens, immune regulation and prevention of autoimmunity (34). Human C-type lectin-like molecule-1 (CLL-1; CLEC12A) is a transmembrane glycoprotein that plays a role in immune regulation as an inhibitory receptor. CLL-1 is present in granulocytes, monocytes and certain types of myeloid progenitors in the bone marrow. Furthermore, CLL-1 is detected in 77.5–92% of AML blasts at diagnosis and is also present in leukemic stem cells, causing treatment failure and leukemia relapse (34). However, the association between the expression of CLL-1 and other classical AML markers remains unclear, and the predictive value of CLL-1 expression in patients with AML has rarely been reported.
SH2 was another representative cluster found in the present study. SH2 domain-containing phosphatase 2 (SHP2; PTPN11) is a positive regulator of receptor tyrosine kinase-driven signaling in response to growth factors and cytokines, including signaling through the Ras/RAF/extracellular signal-regulated kinase (ERK), and the JAK/STAT pathways. Hyperactive SHP2 is associated with tumorigenesis, tumor maintenance, metastasis and therapeutic resistance (35). Numerous somatic gain-of-function mutations that similarly cause the constitutive activation of SHP2 are found in leukemia (35).
In the present study, other clusters observed in the functional analysis were mammalian defensins, which are endogenous peptides produced by certain leukocytes and epithelial cells. In humans, α-defensins are packaged in azurophilic granules of neutrophils or secreted by intestinal Paneth cells. In addition, β-defensins are constitutively expressed in various mucosa and epithelial cells, where they are upregulated in response to infectious and inflammatory stimuli (36). Humans produce six different α-defensins, including four peptides (HNP-1 to HNP-4) in neutrophils and two peptides (HD5 and HD6) in Paneth cells of the small bowel. Several tumor types, including lung, esophageal and skin cancer, exhibit a deregulated expression and secretion of α- and β-defensins (36). The reasons for this deregulated expression and the role of defensins in oncogenesis remain poorly understood.
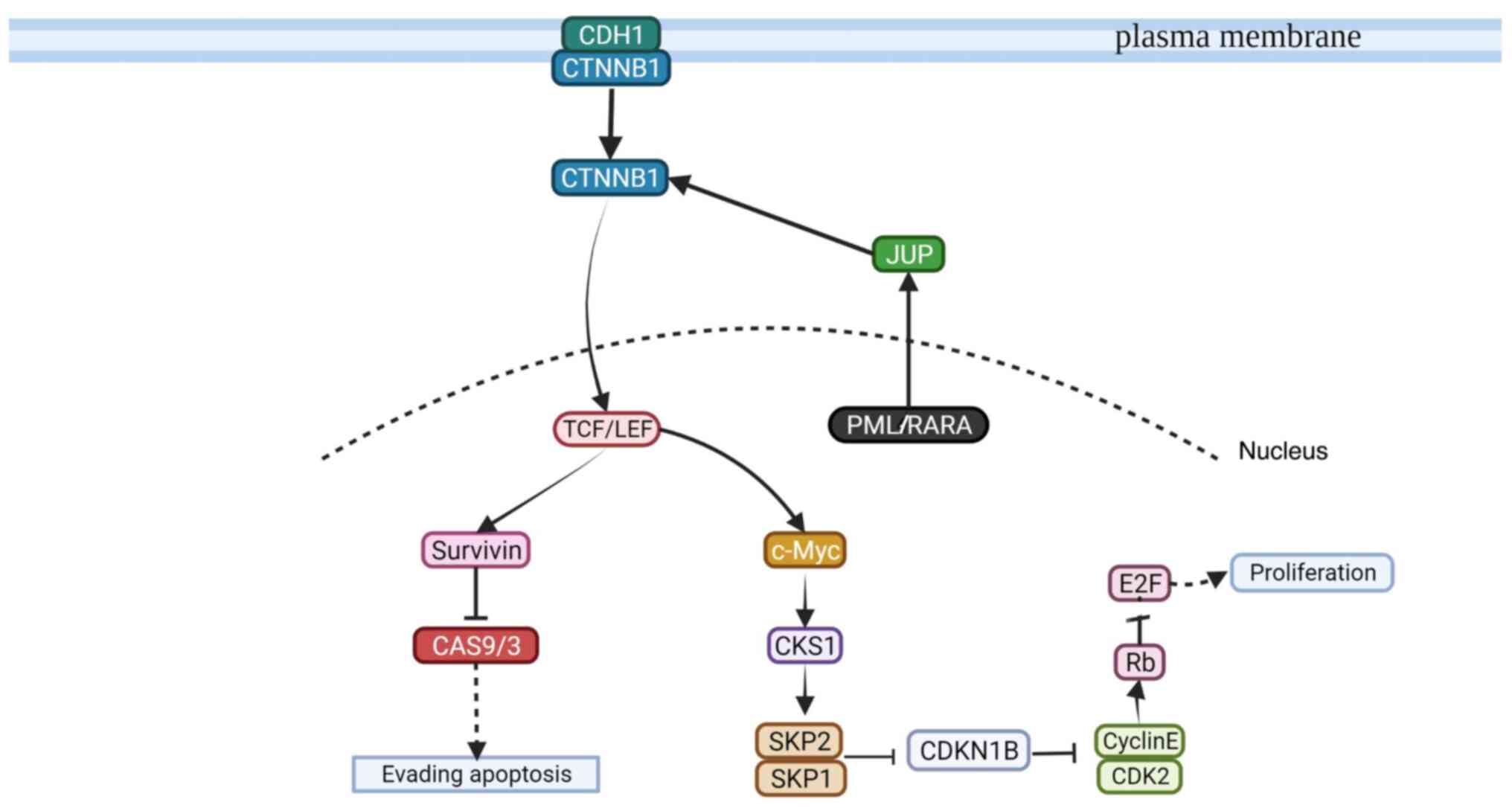
To the best of our knowledge, to date there have been no studies that indicate an association of defensins with AML. Nonetheless, α- and β-defensins have been previously reported to be associated with chronic myeloid leukemia (37–39). However, the present study did not establish an association between PML/RARA transcripts and the functionally identified clusters using gene expression microarray analysis as the identified clusters do not participate in the PML/RARA oncoprotein pathway. It is considered that CCL-1 may emerge as a promising diagnostic biomarker (34). In addition, the HLA-DR class II implication in promyelocytes was negative. Furthermore, the present study suggested a possible signaling pathway involving the PML/RARA oncoprotein that leads to cell proliferation or the evasion of apoptosis, based on the microarray analysis and literature search (Fig. 2).
In conclusion, to the best of our knowledge, the present study is the first to report a pediatric patient with AML with three simultaneous transcripts. The three transcripts may be protective as the patient exhibited a positive response to treatment. As it has been 4 years since the end of the treatment, the patient is considered to be cured. Moreover, the PML/RARA transcripts (bcr1, bcr2 and bcr3) coincide with the good patient prognosis observed in the present case. Further accumulation of data on similar cases may provide relevant clinical evidence for pediatric APL. The functional clusters identified in the patient of the present study may be related to APL biology and suggest promising biomarkers, such as CCL-1, and α- and β-defensins. The PML/RARA oncoprotein signaling pathway suggested in the present study may be associated with the leukemogenic events involved in APL.
Supplementary Material
Supporting Data
Supporting Data
Acknowledgements
The authors would like to thank Dr Violeta Casandra Vera Cuevas (Molecular technical advice; Illumina, Inc.) for support with the genomic analysis.
Funding
The present study was supported by PROSNI (support program to improve the production conditions of research members) 2018 and The Postgraduate Incorporation and Permanence Program from the Postgraduate Human Genetic Department of the University of Guadalajara.
Availability of data and materials
The gene expression datasets generated and/or analyzed during the current study are available in the Gene Expression Omnibus (40) repository (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE205372). This was in-line with the MIAME and MINSEQE guidelines. All other datasets used and/or analyzed during the current study are available from the corresponding author on reasonable request.
Author's contributions
JSH contributed to the design of the study, performed technical procedures, drafted the article and interpreted genomic data. LMM acquired data and performed qPCR procedures. ICQ performed technical microarray procedures. AMM and GSS performed cytogenetics technical procedures. SABJ and RMCO interpreted the clinical data. UFSB contributed to the interpretation and analysis of results. DOS contributed to the interpretation and analysis of microarray results. FASZ coordinated the clinical management. RCR interpreted clinical results and reviewed the manuscript. ACR designed the study and reviewed the submitted version. LBM designed the study and coordinated the final approval of the submitted version. ACR and LBM confirm the authenticity of all the raw data. All authors read and approved the final version of the manuscript.
Ethics approval and consent to participate
The present study was conducted following the principles of The Declaration of Helsinki. The present study was submitted and accepted by The Research Committee and The Research Ethics Committee of The Civil Hospital of Guadalajara (approval no. 00116). Written informed consent and assent were obtained for participation in this study.
Patient consent for publication
Written informed consent was obtained from the patient and their parents to publish the present study.
Competing interests
The authors declare that they have no competing interests.
Glossary
Abbreviations
Abbreviations:
|
APL |
acute promyelocytic leukemia |
|
PML |
promyelocytic leukemia |
|
RARA |
retinoic acid receptor α |
References
|
Testi AM, D'Angiò M, Locatelli F, Pession A and Lo Coco F: Acute promyelocytic leukemia (APL): Comparison between children and adults. Mediterr J Hematol Infect Dis. 6:e20140322014. View Article : Google Scholar : PubMed/NCBI | |
|
Gregory J and Feusner J: Acute promyelocytic leukemia in childhood. Curr Oncol Rep. 11:439–445. 2009. View Article : Google Scholar : PubMed/NCBI | |
|
Jeddi R, Ghédira H, Ben Abdennebi Y, Kacem K, Ben Amor R, Aissaoui L, Bouterâa W, Ben Lakhal R, Ben Abid H, Menif S, et al: ATRA and anthracycline-based chemotherapy in the treatment of childhood acute promyelocytic leukemia (APL): A 10-year experience in Tunisia. Med Oncol. 28:1618–1623. 2011. View Article : Google Scholar : PubMed/NCBI | |
|
Arber DA, Orazi A, Hasserjian R, Thiele J, Borowitz MJ, Le Beau MM, Bloomfield CD, Cazzola M and Vardiman JW: The 2016 revision to the World Health Organization classification of myeloid neoplasms and acute leukemia. Blood. 127:2391–2405. 2016. View Article : Google Scholar : PubMed/NCBI | |
|
Wiernik PH, Gallagher RE and Tallman MS: Acute promyelocytic leukemia. Neoplastic Diseases of the Blood. Springer International Publishing; pp. 409–463. 2018, View Article : Google Scholar | |
|
Cicconi L, Fenaux P, Kantarjian H, Tallman M, Sanz MA and Lo-Coco F: Molecular remission as a therapeutic objective in acute promyelocytic leukemia. Leukemia. 32:1671–1678. 2018. View Article : Google Scholar : PubMed/NCBI | |
|
Kim B, Lee H, Shin S, Lee ST and Choi JR: Clinical evaluation of massively parallel RNA sequencing for detecting recurrent gene fusions in hematologic malignancies. J Mol Diagn. 21:163–170. 2019. View Article : Google Scholar : PubMed/NCBI | |
|
Iaccarino L, Divona M, Ottone T, Cicconi L, Lavorgna S, Ciardi C, Alfonso V, Travaglini S, Facchini L, Cimino G, et al: Identification and monitoring of atypical PML/RARA fusion transcripts in acute promyelocytic leukemia. Genes Chromosomes Cancer. 58:60–65. 2019. View Article : Google Scholar : PubMed/NCBI | |
|
Korf K, Wodrich H, Haschke A, Ocampo C, Harder L, Gieseke F, Pollmann A, Dierck K, Prall S, Staege H, et al: The PML domain of PML-RARα blocks senescence to promote leukemia. Proc Natl Acad Sci USA. 111:12133–12138. 2014. View Article : Google Scholar : PubMed/NCBI | |
|
Conneely SE and Stevens AM: Advances in pediatric acute promyelocytic leukemia. Children. 7:112020. View Article : Google Scholar : PubMed/NCBI | |
|
Liquori A, Ibañez M, Sargas C, Sanz MÁ, Barragán E and Cervera J: Acute promyelocytic leukemia: A constellation of molecular events around a single PML-RARA fusion gene. Cancers (Basel). 12:6242020. View Article : Google Scholar : PubMed/NCBI | |
|
Serdlow SH, Campo E, Pileri SA, Harris NL, Stein H, Siebert R, Advani R, Ghielmini M, Salles GA, Zelenetz AD and Jaffe ES: The 2016 revision of the World Health Organization classification of lymphoid neoplasms. Blood. 2375–2390. 2016. View Article : Google Scholar | |
|
Miguel S: PETHEMA Recomendaciones LPA2012. Hematologialafe. pp1–37. 2012.https://www.sehh.es/servicios-para-los-socios/559-servicios-para-los-socios/pethema/protocolos/lap-2012January 8–2024 | |
|
Rio DC, Ares M, Hannon GJ and Nilsen TW: Purification of RNA using TRIzol (TRI reagent). Cold Spring Harb Protoc. 2010.pdb.prot54392010. View Article : Google Scholar : PubMed/NCBI | |
|
Gabert J, Beillard E, van der Velden VHJ, Bi W, Grimwade D, Pallisgaard N, Barbany G, Cazzaniga G, Cayuela JM, Cavé H, et al: Standardization and quality control studies of ‘real-time’ quantitative reverse transcriptase polymerase chain reaction of fusion gene transcripts for residual disease detection in leukemia-a Europe against cancer program. Leukemia. 17:2318–2357. 2003. View Article : Google Scholar : PubMed/NCBI | |
|
Livak KJ and Schmittgen TD: Analysis of relative gene expression data using real-time quantitative PCR and the 2(−Delta Delta C(T)) method. Methods. 25:402–408. 2001. View Article : Google Scholar : PubMed/NCBI | |
|
Yoo ES: Recent advances in the diagnosis and management of childhood acute promyelocytic leukemia. Korean J Pediatr. 54:95–105. 2011. View Article : Google Scholar : PubMed/NCBI | |
|
Ookura M, Hosono N, Tasaki T, Oiwa K, Fujita K, Ito K, Lee S, Matsuda Y, Morita M, Tai K, et al: Successful treatment of disseminated intravascular coagulation by recombinant human soluble thrombomodulin in patients with acute myeloid leukemia. Medicine (Baltimore). 97:e129812018. View Article : Google Scholar : PubMed/NCBI | |
|
Barragán E, Bolufer P, Martín G, Cervera J, Moreno I, Capote FJ, Rosique P and Sanz MA: Identification of two atypical PML-RAR(alpha) transcripts in two patients with acute promyelocytic leukemia. Leuk Res. 26:439–442. 2002. View Article : Google Scholar : PubMed/NCBI | |
|
Kim M, Lim J, Kim Y, Han K, Lee DH, Chung NG, Cho B, Kim HK, Eom KS, Min CK and Min WS: The genetic characterization of acute promyelocytic leukemia with cryptic t(15;17) including a new recurrent additional cytogenetic abnormality i(17)(q10). Leukemia. 22:881–883. 2008. View Article : Google Scholar : PubMed/NCBI | |
|
Kim MJ, Cho SY, Kim MH, Lee JJ, Kang SY, Cho EH, Huh J, Yoon HJ, Park TS, Lee WI, et al: FISH-negative cryptic PML-RARA rearrangement detected by long-distance polymerase chain reaction and sequencing analyses: A case study and review of the literature. Cancer Genet Cytogenet. 203:278–283. 2010. View Article : Google Scholar : PubMed/NCBI | |
|
Manola KN, Karakosta M, Sambani C, Terzoudi G, Pagoni M, Gatsa E and Papaioannou M: Isochromosome der(17)(q10)t(15;17) in acute promyelocytic leukemia resulting in an additional copy of the RARA-PML fusion gene: Report of 4 cases and review of the literature. Acta Haematol. 123:162–170. 2010. View Article : Google Scholar : PubMed/NCBI | |
|
Rasekh EO, Elsayed GM, Madney Y and El Gammal MM: Prognostic significance of bcr-1 and bcr-3 isoforms of PML-RARA and FLT3-ITD in patients with acute promyelocytic leukemia. Clin Lymphoma Myeloma Leuk. 20:156–167. 2020. View Article : Google Scholar : PubMed/NCBI | |
|
Rasekh EO, Elsayed GM and Fathy S: No prognostic significance of normalized copy number of PML-RARA transcript at diagnosis in patients with acute promyelocytic leukemia. Hematol Oncol Stem Cell Ther. 14:119–125. 2021. View Article : Google Scholar : PubMed/NCBI | |
|
Heyer EE, Deveson IW, Wooi D, Selinger CI, Lyons RJ, Hayes VM, O'Toole SA, Ballinger ML, Gill D, Thomas DM, et al: Diagnosis of fusion genes using targeted RNA sequencing. Nat Commun. 10:13882019. View Article : Google Scholar : PubMed/NCBI | |
|
Loeb LA: A mutator phenotype in cancer. Cancer Res. 61:3230–3239. 2001.PubMed/NCBI | |
|
De Braekeleer E, Douet-Guilbert N and De Braekeleer M: RARA fusion genes in acute promyelocytic leukemia: A review. Expert Rev Hematol. 7:347–357. 2014. View Article : Google Scholar : PubMed/NCBI | |
|
Castro-Mujica Mdel C and Sullcahuamán-Allende Y: Molecular subtypes of PML/RARα in patients with acute promyelocytic leukemia. Rev Peru Med Exp Salud Publica. 30:37–40. 2013.(In Spanish). PubMed/NCBI | |
|
Christopher MJ, Petti AA, Rettig MP, Miller CA, Chendamarai E, Duncavage EJ, Klco JM, Helton NM, O'Laughlin M, Fronick CC, et al: Immune escape of relapsed AML cells after allogeneic transplantation. N Engl J Med. 379:2330–2341. 2018. View Article : Google Scholar : PubMed/NCBI | |
|
Dunn GP, Bruce AT, Ikeda H, Old LJ and Schreiber RD: Cancer immunoediting: From immunosurveillance to tumor escape. Nat Immunol. 3:991–998. 2002. View Article : Google Scholar : PubMed/NCBI | |
|
Oelschlaegel U, Mohr B, Schaich M, Schäkel U, Kroschinsky F, Illmer T, Ehninger G and Thiede C: HLA-DRneg patients without acute promyelocytic leukemia show distinct immunophenotypic, genetic, molecular, and cytomorphologic characteristics compared to acute promyelocytic leukemia. Cytometry B Clin Cytom. 76:321–327. 2009. View Article : Google Scholar : PubMed/NCBI | |
|
Roerden M, Märklin M, Salih HR, Bethge WA, Klein R, Rammensee HG, Nelde A and Walz JS: Expression levels of HLA-DR in acute myeloid leukemia: Implications for antigenicity and clinical outcome. Leuk Lymphoma. 62:1907–1919. 2021. View Article : Google Scholar : PubMed/NCBI | |
|
Nie L, Ma R, Yuan X, Jiang L, Yang S, Xu H, Liu X, Liu Y, Zhang L and Zhu Z: The prognostic value of CD2, CD4, and HLA-DR expression and FLT3-ITD mutation in adult acute promyelocytic leukemia. Leuk Lymphoma. 61:2482–2487. 2020. View Article : Google Scholar : PubMed/NCBI | |
|
Wang J, Wang W, Chen H, Li W, Huang T, Zhang W, Ling W, Lai P, Wang Y, Geng S, et al: C-type lectin-like molecule-1 as a biomarker for diagnosis and prognosis in acute myeloid leukemia: A preliminary study. Biomed Res Int. 2021:66439482021.PubMed/NCBI | |
|
Raveendra-Panickar D, Finlay D, Layng FI, Lambert LJ, Celeridad M, Zhao M, Barbosa K, De Backer LJS, Kwong E, Gosalia P, et al: Discovery of novel furanylbenzamide inhibitors that target oncogenic tyrosine phosphatase SHP2 in leukemia cells. J Biol Chem. 298:1014772022. View Article : Google Scholar : PubMed/NCBI | |
|
Ghosh SK, McCormick TS and Weinberg A: Human beta defensins and cancer: Contradictions and common ground. Front Oncol. 9:3412019. View Article : Google Scholar : PubMed/NCBI | |
|
Stretch C, Khan S, Asgarian N, Eisner R, Vaisipour S, Damaraju S, Graham K, Bathe OF, Steed H, Greiner R and Baracos VE: Effects of sample size on differential gene expression, rank order and prediction accuracy of a gene signature. PLoS One. 8:e653802013. View Article : Google Scholar : PubMed/NCBI | |
|
Maleki F, Ovens K, McQuillan I and Kusalik AJ: Size matters: How sample size affects the reproducibility and specificity of gene set analysis. Hum Genomics. 13 (Suppl 1):S422019. View Article : Google Scholar : PubMed/NCBI | |
|
Tsai CA, Wang SJ, Chen DT and Chen JJ: Sample size for gene expression microarray experiments. Bioinformatics. 21:1502–1508. 2005. View Article : Google Scholar : PubMed/NCBI | |
|
Edgar R, Domrachev M and Lash AE: Gene expression omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 30:207–210. 2002. View Article : Google Scholar : PubMed/NCBI | |
|
Kent WJ, Sugnet CW, Furey TS, Roskin KM, Pringle TH, Zahler AM and Haussler D: The human genome browser at UCSC. Genome Res. 12:996–1006. 2002. View Article : Google Scholar : PubMed/NCBI |