|
1
|
Trayhurn P: Of genes and genomes- and dark
matter. Br J Nutr. 91:1–2. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Palazzo AF and Lee ES: Non-coding RNA:
What is functional and what is junk? Front Genet. 6:22015.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Pennisi E: Shining a light on the genome's
'Dark Matter.'. Science. 330:16142010. View Article : Google Scholar
|
|
4
|
Amaral PP and Mattick JS: Noncoding RNA in
development. Mamm Genome. 19:454–492. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Jacob F and Monod J: Genetic regulatory
mechanisms in the synthesis of proteins. J Mol Biol. 3:318–356.
1961. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Nerenz RD and Lefferts J: Our genome's
'Dark Matter' is the next frontier in molecular diagnostics. Clin
Chem. 63:792–793. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Qureshi IA, Mattick JS and Mehler MF: Long
non-coding RNAs in nervous system function and disease. Brain Res.
1338:20–35. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Salviano-Silva A, Lobo-Alves SC, Almeida
RC, Malheiros D and Petzl-Erler ML: Besides pathology: Long
Non-Coding RNA in cell and tissue homeostasis. Noncoding RNA.
4:32018. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
de Kloet ER, Fitzsimons CP, Datson NA,
Meijer OC and Vreugdenhil E: Glucocorticoid signaling and
stress-related limbic susceptibility pathway: About receptors,
transcription machinery and microRNA. Brain Res. 1293:129–141.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
de Almeida RA, Fraczek MG, Parker S,
Delneri D and O'Keefe RT: Non-coding RNAs and disease: The
classical ncRNAs make a comeback. Biochem Soc Trans. 44:1073–1078.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Kaikkonen MU, Lam MT and Glass CK:
Non-coding RNAs as regulators of gene expression and epigenetics.
Cardiovasc Res. 90:430–440. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Babenko O, Golubov A, Ilnytskyy Y,
Kovalchuk I and Metz GA: Genomic and epigenomic responses to
chronic stress involve miRNA-mediated programming. PLoS One.
7:e294412012. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Rinaldi A, Vincenti S, De Vito F, Bozzoni
I, Oliverio A, Presutti C, Fragapane P and Mele A: Stress induces
region specific alterations in microRNAs expression in mice. Behav
Brain Res. 208:265–269. 2010. View Article : Google Scholar
|
|
14
|
Daskalakis NP, Provost AC, Hunter RG and
Guffanti G: Noncoding RNAs: Stress, glucocorticoids, and
posttraumatic stress disorder. Biol Psychiatry. 83:849–865. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Hombach S and Kretz M: Non-coding RNAs:
Classification, biology and functioning. Adv Exp Med Biol.
937:3–17. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Mattick JS and Makunin IV: Small
regulatory RNAs in mammals. Hum Mol Genet. 14(Spec No 1):
R121–R132. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Lam JK, Chow MY, Zhang Y and Leung SW:
siRNA Versus miRNA as therapeutics for gene silencing. Mol Ther
Nucleic Acids. 4:e2522015. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Harrow J, Frankish A, Gonzalez JM,
Tapanari E, Diekhans M, Kokocinski F, Aken BL, Barrell D, Zadissa
A, Searle S, et al: GENCODE: The reference human genome annotation
for The ENCODE Project. Genome Res. 22:1760–1774. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Pennacchio LA, Bickmore W, Dean A, Nobrega
MA and Bejerano G: Enhancers: Five essential questions. Nat Rev
Genet. 14:288–295. 2013. View
Article : Google Scholar : PubMed/NCBI
|
|
20
|
Jo BS and Choi SS: Introns: The functional
benefits of introns in genomes. Genomics Inform. 13:112–118. 2015.
View Article : Google Scholar
|
|
21
|
Mattick JS and Makunin IV: Non-coding RNA.
Hum Mol Genet. 15(Spec No 1): R17–R29. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Wahid F, Shehzad A, Khan T and Kim YY:
MicroRNAs: Synthesis, mechanism, function, and recent clinical
trials. Biochim Biophys Acta. 1803:1231–1243. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Croce CM and Calin GA: miRNAs, cancer, and
stem cell division. Cell. 122:6–7. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Maggert KA: Stress: An evolutionary
mutagen. Proc Natl Acad Sci USA. 116:17616–17618. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Wheeler BS: Small RNAs, big impact: Small
RNA pathways in transposon control and their effect on the host
stress response. Chromosome Res. 21:587–600. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Wang KC and Chang HY: Molecular mechanisms
of long noncoding RNAs. Mol Cell. 43:904–914. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Amaral PP, Dinger ME and Mattick JS:
Non-coding RNAs in homeostasis, disease and stress responses: An
evolutionary perspective. Brief Funct Genomics. 12:254–278. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Cipolla GA, de Oliveira JC, Salviano-Silva
A, Lobo-Alves SC, Lemos DS, Oliveira LC, Jucoski TS, Mathias C,
Pedroso GA, Zambalde EP and Gradia DF: Long Non-Coding RNAs in
multifactorial diseases: Another layer of complexity. Noncoding
RNA. 4:132018. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Kung JT, Colognori D and Lee JT: Long
noncoding RNAs: Past, present, and future. Genetics. 193:651–669.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
de Lara JC, Arzate-Mejía RG and
Recillas-Targa F: Enhancer RNAs: Insights into their biological
role. Epigenet Insights. 12:25168657198460932019. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Wu M and Shen J: From super-enhancer
Non-coding RNA to immune checkpoint: Frameworks to functions. Front
Oncol. 9:13072019. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Chen H, Du G, Song X and Li L: Non-coding
transcripts from enhancers: New insights into enhancer activity and
gene expression regulation. Genomics Proteomics Bioinformatics.
15:201–207. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Parenteau J and Abou Elela S: Introns:
Good day junk is bad day treasure. Trends Genet. 35:923–934. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Hubé F and Francastel C: Mammalian
introns: When the junk generates molecular diversity. Int J Mol
Sci. 16:4429–4452. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Pink RC, Wicks K, Caley DP, Punch EK,
Jacobs L and Carter DR: Pseudogenes: Pseudo-functional or key
regulators in health and disease? RNA. 17:792–798. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Oliva-Rico D and Herrera LA: Regulated
expression of the lncRNA TERRA and its impact on telomere biology.
Mech Ageing Dev. 167:16–23. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Luke B and Lingner J: TERRA: Telomeric
repeat-containing RNA. EMBO J. 28:2503–2510. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Schmidt U, Keck ME and Buell DR: miRNAs
and other non-coding RNAs in posttraumatic stress disorder: A
systematic review of clinical and animal studies. J Psychiatr Res.
65:1–8. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Storey KB and Wu CW: Stress response and
adaptation: A new molecular toolkit for the 21st entury. Comp
Biochem Physiol. Part A Mol Integr Physiol. 165:417–428. 2013.
View Article : Google Scholar
|
|
40
|
Malan-Müller S, Hemmings SM and Seedat S:
Big effects of small RNAs: A review of MicroRNAs in Anxiety. Mol
Neurobiol. 47:726–739. 2013. View Article : Google Scholar :
|
|
41
|
Hollins SL and Cairns MJ: MicroRNA: Small
RNA mediators of the brains genomic response to environmental
stress. Prog Neurobiol. 143:61–81. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Nemoto T and Kakinuma Y: Involvement of
Noncoding RNAs in stress-related neuropsychiatric diseases caused
by DOHaD Theory: ncRNAs and DOHaD-Induced neuropsychiatric
diseases. Adv Exp Med Biol. 1012:49–59. 2018. View Article : Google Scholar
|
|
43
|
Levran O, Correa da Rosa J, Randesi M,
Rotrosen J, Adelson M and Kreek MJ: A non-coding CRHR2 SNP
rs255105, a cis-eQTL for a downstream lincRNA AC005154.6, is
associated with heroin addiction. PLoS One. 13:e01999512018.
View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Kim KW: PIWI Proteins and piRNAs in the
Nervous System. Mol Cells. 42:828–835. 2019.PubMed/NCBI
|
|
45
|
Gapp K, Jawaid A, Sarkies P, Bohacek J,
Pelczar P, Prados J, Farinelli L, Miska E and Mansuy IM:
Implication of sperm RNAs in transgenerational inheritance of the
effects of early trauma in mice. Nat Neurosci. 17:667–669. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Bystritsky A, Khalsa SS, Cameron ME and
Schiffman J: Current diagnosis and treatment of anxiety disorders.
P T. 38:30–57. 2013.PubMed/NCBI
|
|
47
|
Schouten M, Aschrafi A, Bielefeld P,
Doxakis E and Fitzsimons CP: microRNAs and the regulation of
neuronal plasticity under stress conditions. Neuroscience.
241:188–205. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Li YJ, Xu M, Gao ZH, Wang YQ, Yue Z, Zhang
YX, Li XX, Zhang C, Xie SY and Wang PY: Alterations of serum levels
of BDNF-related miRNAs in patients with depression. PLoS One.
8:e636482013. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
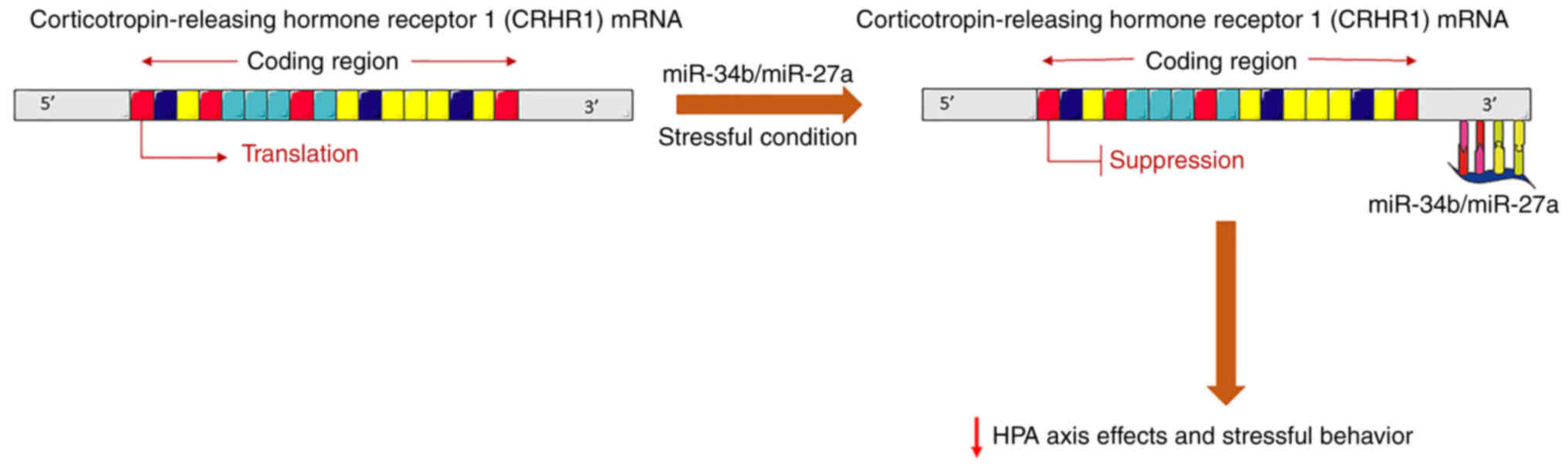
Zhu J, Chen Z, Tian J, Meng Z, Ju M, Wu G
and Tian Z: miR-34b attenuates trauma-induced anxiety-like behavior
by targeting CRHR1. Int J Mol Med. 40:90–100. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Volk N, Pape JC, Engel M, Zannas AS,
Cattane N, Cattaneo A, Binder EB and Chen A: Amygdalar MicroRNA-15a
is essential for coping with chronic stress. Cell Rep.
17:1882–1891. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
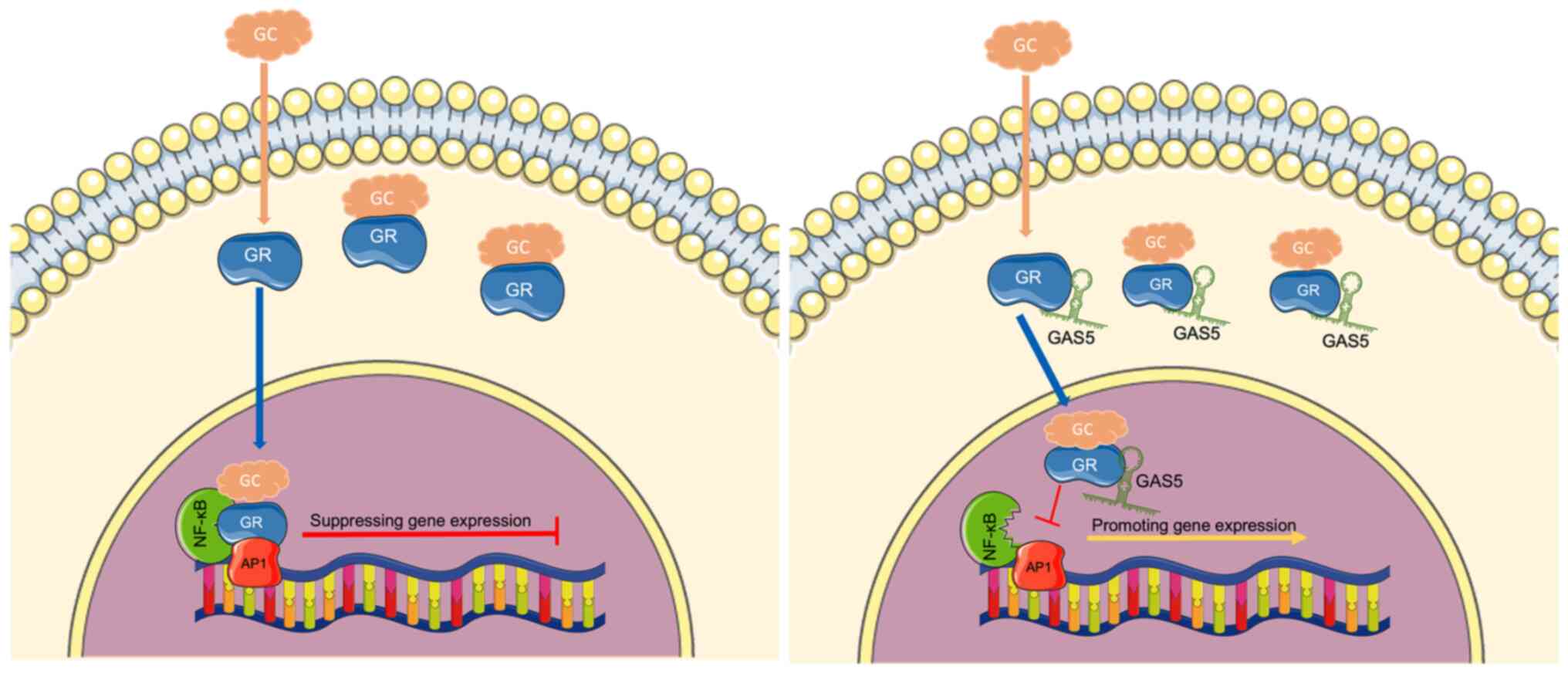
Kino T, Hurt DE, Ichijo T, Nader N and
Chrousos GP: Noncoding RNA Gas5 is a growth arrest and
starvation-associated repressor of the glucocorticoid receptor. Sci
Signal. 3:ra82010. View Article : Google Scholar :
|
|
52
|
Mayama T, Marr AK and Kino T: Differential
expression of glucocorticoid receptor Noncoding RNA repressor Gas5
in autoimmune and inflammatory diseases. Horm Metab Res.
48:550–557. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Moradi M, Gharesouran J, Ghafouri-Fard S,
Noroozi R, Talebian S, Taheri M and Rezazadeh M: Role of NR3C1 and
GAS5 genes polymorphisms in multiple sclerosis. Int J Neurosci.
130:407–412. 2020. View Article : Google Scholar
|
|
54
|
Lucafò M, Bravin V, Tommasini A,
Martelossi S, Rabach I, Ventura A, Decorti G and De Iudicibus S:
Differential expression of GAS5 in rapamycin-induced reversion of
glucocorticoid resistance. Clin Exp Pharmacol Physiol. 43:602–605.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Kukharsky MS, Ninkina NN, An H, Telezhkin
V, Wei W, Meritens CR, Cooper-Knock J, Nakagawa S, Hirose T,
Buchman VL and Shelkovnikova TA: Long non-coding RNA Neat1
regulates adaptive behavioural response to stress in mice. Transl
Psychiatry. 10:1712020. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Tilstra JS, Clauson CL, Niedernhofer LJ
and Robbins PD: NF-κB in aging and disease. Aging Dis. 2:449–465.
2011.
|
|
57
|
Rapicavoli NA, Qu K, Zhang J, Mikhail M,
Laberge RM and Chang HY: A mammalian pseudogene lncRNA at the
interface of inflammation and anti-inflammatory therapeutics.
ELife. 2:e007622013. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Mai C, Qiu L, Zeng Y and Jian HG: LncRNA
Lethe protects sepsis-induced brain injury via regulating autophagy
of cortical neurons. Eur Rev Med Pharmacol Sci. 23:4858–4864.
2019.PubMed/NCBI
|
|
59
|
Posner R, Toker IA, Antonova O, Star E,
Anava S, Azmon E, Hendricks M, Bracha S, Gingold H and Rechavi O:
Neuronal small RNAs control behavior transgenerationally. Cell.
177:1814–1826.e15. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Kuintzle RC, Chow ES, Westby TN, Gvakharia
BO, Giebultowicz JM and Hendrix DA: Circadian deep sequencing
reveals stress-response genes that adopt robust rhythmic expression
during aging. Nat Commun. 8:145292017. View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Minerath RA, Hall DD and Grueter CE:
Targeting transcriptional machinery to inhibit enhancer-driven gene
expression in heart failure. Heart Fail Rev. 24:725–741. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Mirtschink P, Bischof C, Pham MD, Sharma
R, Khadayate S, Rossi G, Fankhauser N, Traub S, Sossalla S, Hagag
E, et al: Inhibition of the hypoxia-inducible factor 1α-induced
cardio-specific HERNA1 enhance-templated RNA protects from heart
disease. Circulation. 139:2778–2792. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Aguilo F, Li S, Balasubramaniyan N, Sancho
A, Benko S, Zhang F, Vashisht A, Rengasamy M, Andino B, Chen CH, et
al: Deposition of 5-Methylcytosine on enhancer RNAs enables the
coactivator function of PGC-1α. Cell Rep. 14:479–492. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Sasse SK, Gruca M, Allen MA, Kadiyala V,
Song T, Gally F, Gupta A, Pufall MA, Dowell RD and Gerber AN:
Nascent transcript analysis of glucocorticoid crosstalk with TNF
defines primary and cooperative inflammatory repression. Genome
Res. 29:1753–1765. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
65
|
IIott NE, Heward JA, Roux B, Tsitsiou E,
Fenwick PS, Lenzi L, Goodhead I, Hertz-Fowler C, Heger A, Hall N,
et al: Long non-coding RNAs and enhancer RNAs regulate the
lipopolysaccharide-induced inflammatory response in human
monocytes. Nat Commun. 5:39792014. View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Hanna J, Hossain GS and Kocerha J: The
potential for microRNA therapeutics and clinical research. Front
Genet. 10:4782019. View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Peedicayil J: The potential role of
epigenetic drugs in the treatment of anxiety disorders.
Neuropsychiatr Dis Treat. 16:597–606. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Katsuura S, Kuwano Y, Yamagishi N,
Kurokawa K, Kajita K, Akaike Y, Nishida K, Masuda K, Tanahashi T
and Rokutan K: MicroRNAs miR-144/144* and miR-16 in peripheral
blood are potential biomarkers for naturalistic stress in healthy
Japanese medical students. Neurosci Lett. 516:79–84. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
69
|
Scott KA, Hoban AE, Clarke G, Moloney GM,
Dinan TG and Cryan JF: Thinking small: Towards microRNA-based
therapeutics for anxiety disorders. Expert Opin Investig Drugs.
24:529–542. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
70
|
Moldovan L, Batte KE, Trgovcich J, Wisler
J, Marsh CB and Piper M: Methodological challenges in utilizing
miRNAs as circulating biomarkers. J Cell Mol Med. 18:371–390. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
71
|
Anfossi S, Babayan A, Pantel K and Calin
GA: Clinical utility of circulating non-coding RNAs-an update. Nat
Rev Clin Oncol. 15:541–563. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
72
|
Pascale E, Divisato G, Palladino R,
Auriemma M, Ngalya EF and Caiazzo M: Noncoding RNAs and midbrain DA
neurons: Novel molecular mechanisms and therapeutic targets in
health and disease. Biomolecules. 10:12692020. View Article : Google Scholar : PubMed/NCBI
|
|
73
|
Varela MA, Roberts TC and Wood MJA:
Epigenetics and ncRNAs in brain function and disease: Mechanisms
and prospects for therapy. Neurotherapeutics. 10:621–631. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
74
|
Juliano RL: The delivery of therapeutic
oligonucleotides. Nucleic Acids Res. 44:6518–6548. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
75
|
O'Neill CP and Dwyer RM:
Nanoparticle-Based delivery of tumor suppressor microRNA for cancer
therapy. Cells. 9:5212020. View Article : Google Scholar : PubMed/NCBI
|